Variants
The Variants preference is used to record a list of variants in the system. These are found when adding a gene mutation in the Diagnosis tab. The choice of values are selected using an autocomplete or dropdown list.
Location
The table of Variants will display the code, name, gene, and status for the variant.
Configuration
When adding or editing a variant, users are presented with the below form to complete. Always press Save in the bottom right-hand corner of the browser after adding or editing a variant.
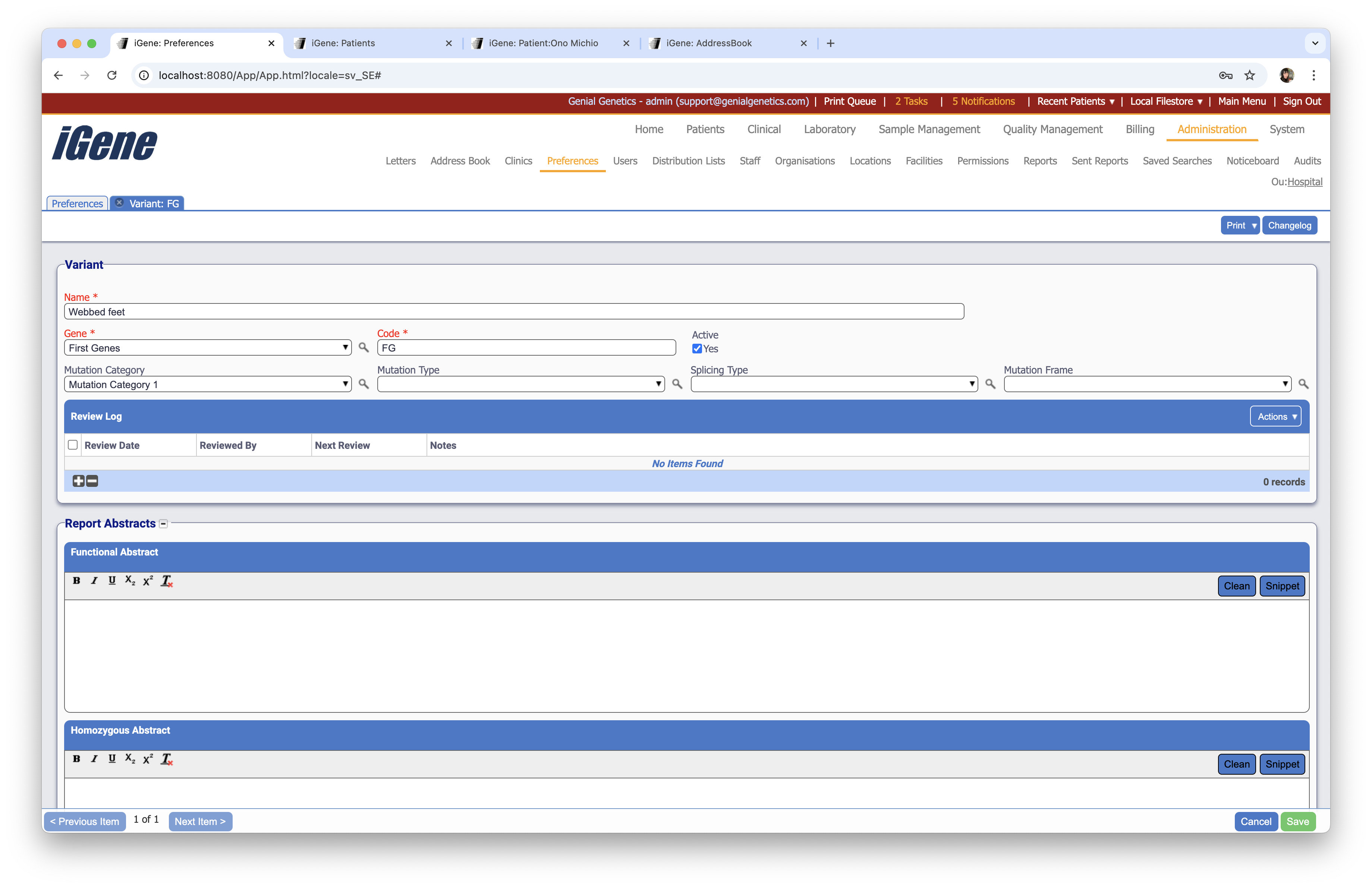
Name (Required)
A name for the variant. It is advisable to keep this unique.
Gene (Required)
A dropdown list of genes to assign to the variant. This is set by the Gene preference.
Code (Required, Unique)
A unique code for the variant. This can be the same as the name but must be unique throughout the entire system.
Active
A checkbox to activate/deactivate an entry. If the preference is not active, it will not be selectable in any drop-down lists.
Mutation Category
A dropdown list of mutation categories to assign to the variant. This is set by the Mutation Categories preference.
Mutation Type
A dropdown list of mutation types to assign to the variant. This is set by the Mutation Types preference.
Splicing Type
A dropdown list of splicing types to assign to the variant. This is set by the Splicing Types preference.
Mutation Frame
A dropdown list of mutation frames to assign to the variant. This is set by the Mutation Frames preference.
Review Log
A table to log reviews of the variant.
Review Date
The date the review was made.
Reviewed By
The user who reviewed the variant.
Next Review
The date the variant is next due for review.
Notes
Any notes made during the review.
Report Abstracts
Functional Abstract
A text field to enter a functional abstract for the variant.
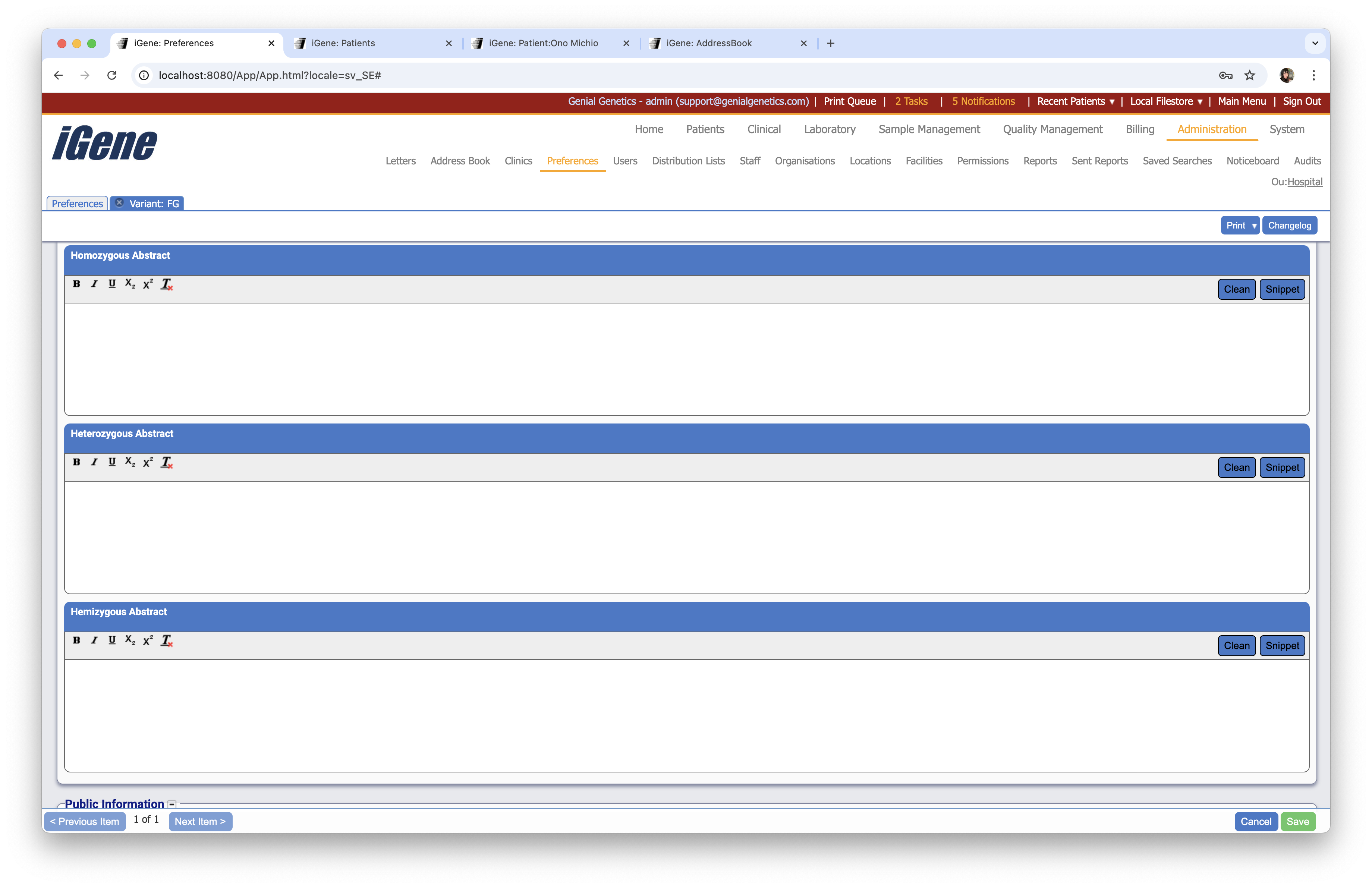
Homozygous Abstract
A text field to enter a homozygous abstract for the variant.
Heterozygous Abstract
A text field to enter a heterozygous abstract for the variant.
Hemizygous Abstract
A text field to enter a hemizygous abstract for the variant.
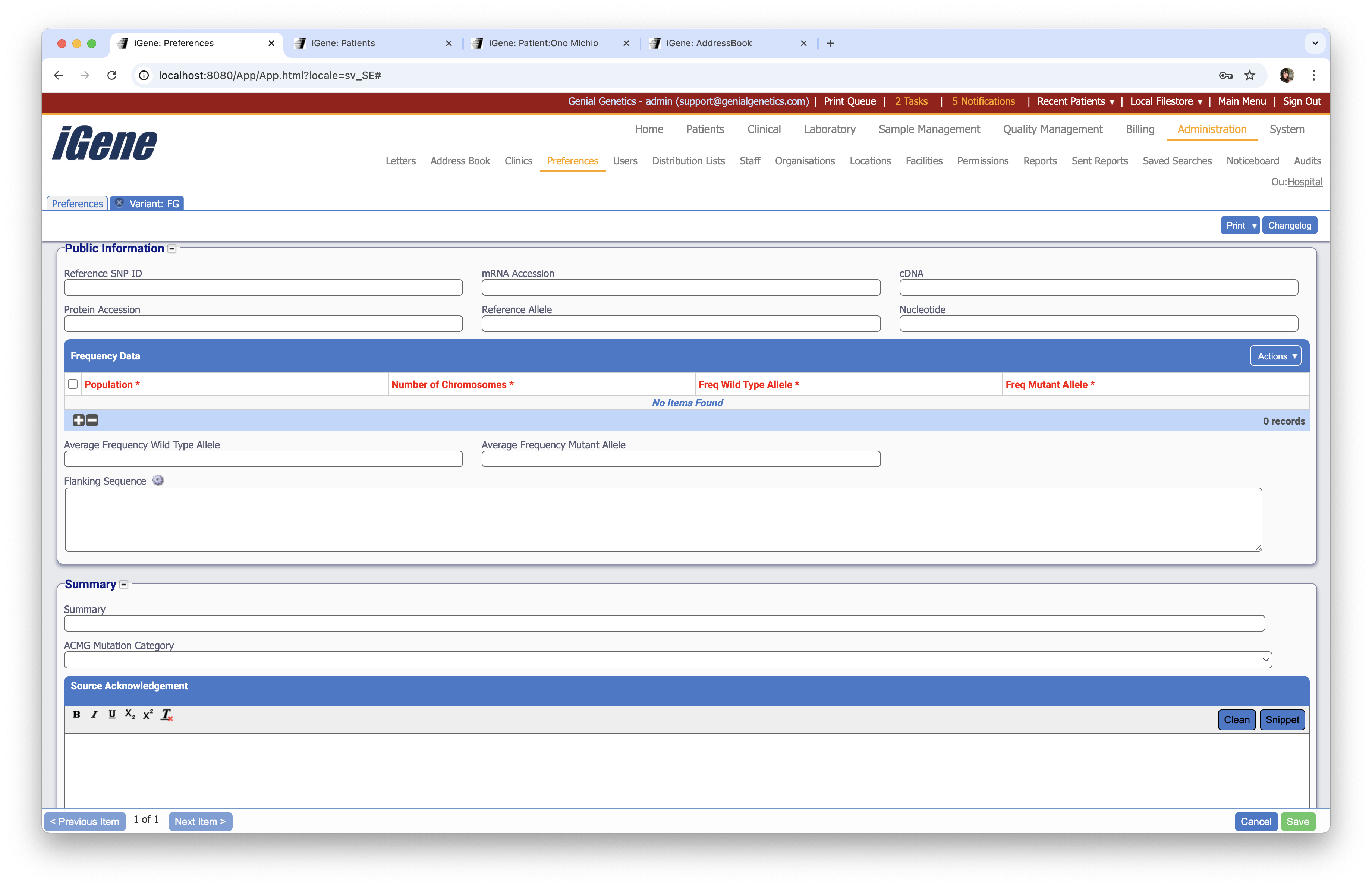
Public Information
Reference SNP ID
A text field to enter the reference SNP ID for the variant.
mRNA Accession
A text field to enter the mRNA accession for the variant.
cDNA
A text field to enter the cDNA for the variant.
Protein Accession
A text field to enter the protein accession for the variant.
Reference Allele
A text field to enter the reference allele for the variant.
Nucleotide
A text field to enter the nucleotide for the variant.
Frequency Data
A table to enter frequency data for the variant.
Population
A text field to enter the population for the frequency data.
Number of Chromosomes
A text field to enter the number of chromosomes for the frequency data.
Freq Wild Type Allele
A text field to enter the frequency of the wild type allele.
Freq Mutant Allele
A text field to enter the frequency of the mutant allele.
Average Frequency Wild Type Allele
A text field to enter the average frequency of the wild type allele.
Average Frequency Mutant Allele
A text field to enter the average frequency of the mutant allele.
Flanking Sequence
A text field to enter the flanking sequence for the variant.
Summary
Summary
A text field to enter a summary for the variant.
ACMG Mutation Category
A dropdown list of ACMG mutation categories to assign to the variant. This is set by the ACMG Mutation Categories preference.
Source Acknowledgement
A text field to enter the source acknowledgement for the variant.
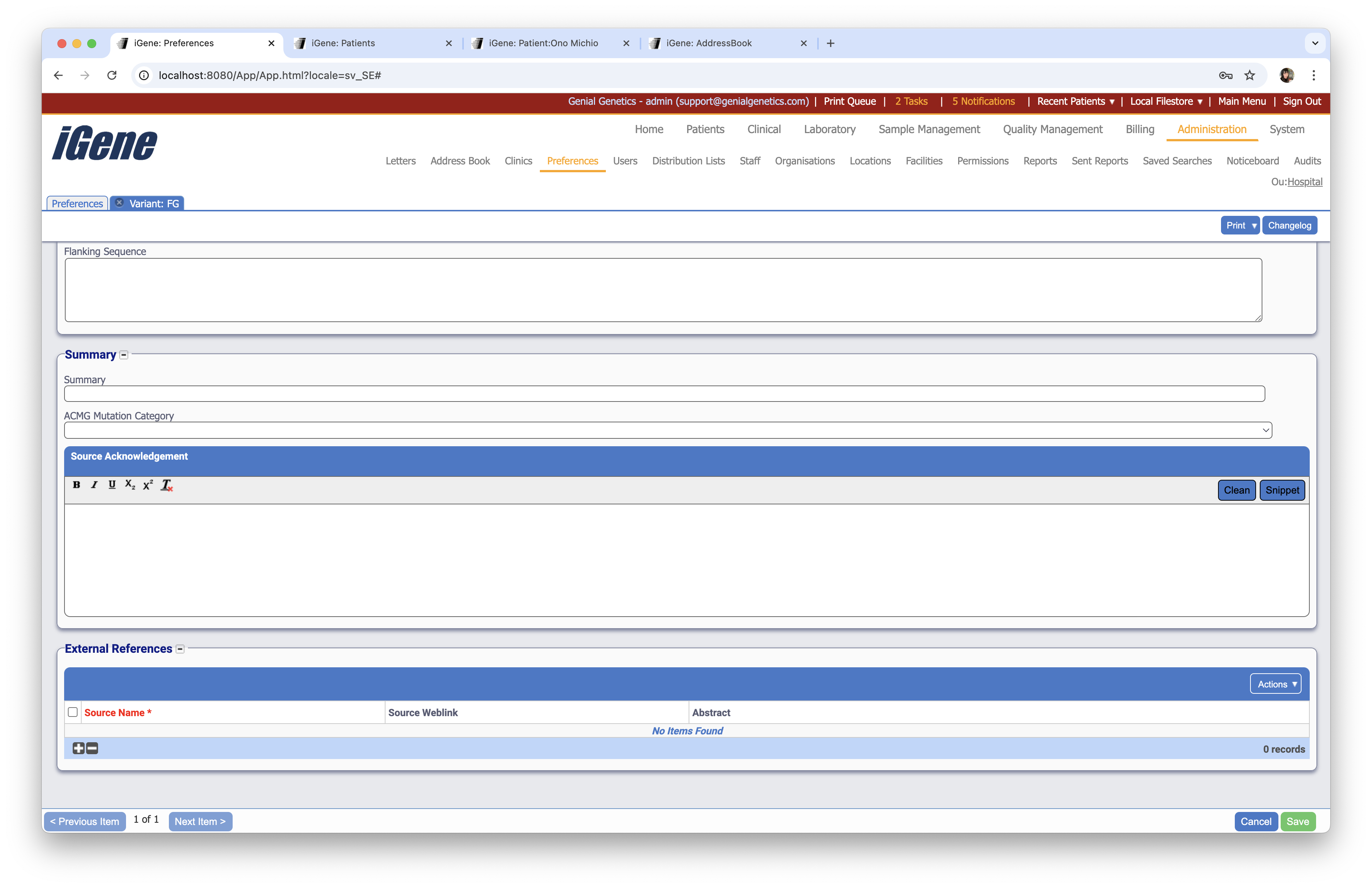
External References
A table to enter external references for the variant.
Source Name
A text field to enter the source name for the external reference.
Source Weblink
A text field to enter the source weblink for the external reference.
Abstract
A text field to enter the abstract for the external reference.